Brainnetome Atlas

Lingzhong Fan 1, Hai Li 1,2, Junjie Zhuo 4, Yu Zhang 1,2, Jiaojian Wang 4, Liangfu Chen 1,2, Zhengyi Yang 1,2, Congying Chu 1,2, Sangma Xie 1,2, Angela R. Laird 10, Peter T. Fox 7, Simon B. Eickhoff 8,9, Chunshui Yu 6 and Tianzi Jiang 1,2,3,4,5
Author information:
1Brainnetome Center, 2National Laboratory of Pattern Recognition and, 3CAS Center for Excellence in Brain Science and Intelligence Technology, Institute of Automation, Chinese Academy of Sciences, Beijing 100190, China, 4Key Laboratory for NeuroInformation of the Ministry of Education, School of Life Science and Technology, University of Electronic Science and Technology of China, Chengdu 625014, China, 5The Queensland Brain Institute, University of Queensland, Brisbane, QLD 4072, Australia, 6Department of Radiology, Tianjin Medical University General Hospital, Tianjin, China, 7Research Imaging Institute, University of Texas Health Science Center, San Antonio, TX, USA, 8Institute of Neuroscience and Medicine (INM-1), Research Centre Juelich, Juelich 52425, Germany, 9Institute for Clinical Neuroscience and Medical Psychology, Heinrich-Heine-University Düsseldorf, Düsseldorf 40225, Germany and 10Department of Physics, Florida International University, Miami, FL, USA
Lingzhong Fan and Hai Li equally contributed to this work.
Abstract:
The human brain atlases that allowcorrelating brain anatomy with psychological and cognitive functions are in transition from ex vivo histology-based printed atlases to digital brain maps providing multimodal in vivo information. Many current human brain atlases cover only specific structures, lack fine-grained parcellations, and fail to provide functionally important connectivity information. Using noninvasive multimodal neuroimaging techniques, we designed a connectivity-based parcellation framework that identifies the subdivisions of the entire human brain, revealing the in vivo connectivity architecture. The resulting human Brainnetome Atlas, with 210 cortical and 36 subcortical subregions, provides a fine-grained, cross-validated atlas and contains information on both anatomical and functional connections. Additionally, we further mapped the delineated structures to mental processes by reference to the BrainMap database. It thus provides an objective and stable starting point from which to explore the complex relationships between structure, connectivity, and function, and eventually improves understanding of howthe human brain works. The human Brainnetome Atlas will be made freely available for download at http://atlas.brainnetome.org, so that whole brain parcellations, connections, and functional data will be readily available for researchers to use in their investigations into healthy and pathological states.
Keywords: brain atlas, connectivity-based parcellation, diffusion tensor imaging, functional characterization, resting-state functional connectivity
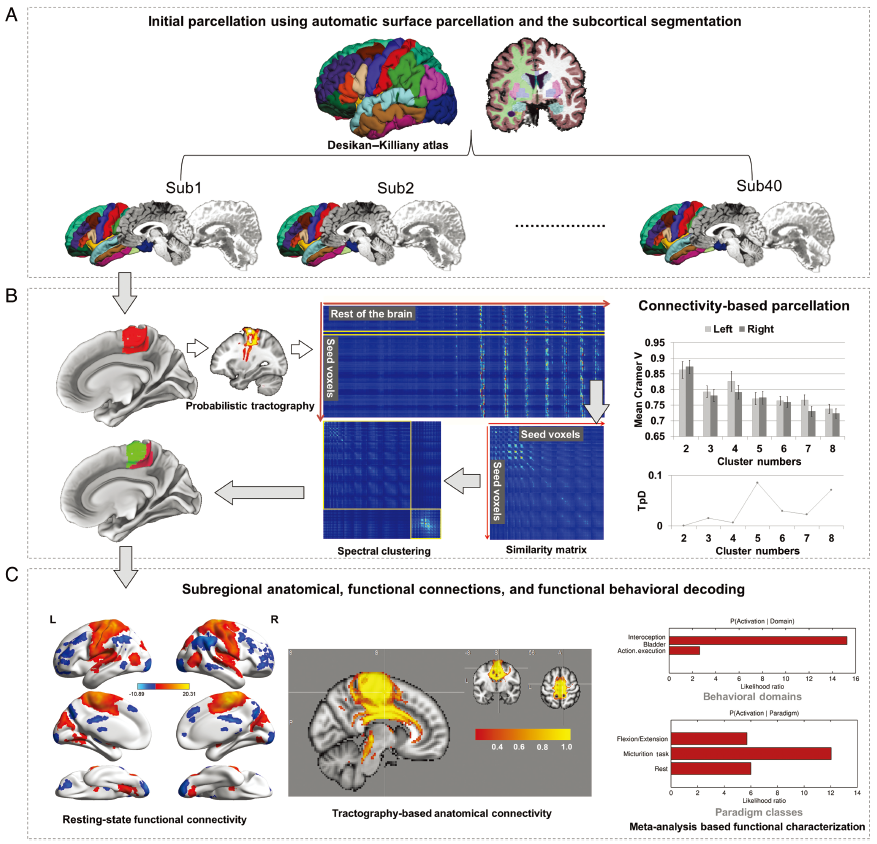
Figure 1. Framework of the Brainnetome Atlas construction based on connectivity-based parcellation. (A) Initial parcellation using automatic surface parcellation and subcortical segmentation. The FreeSurfer DK atlas produced the initial parcellations based on gyri and sulci. (B) Tractography-based parcellation with in vivo connectional architecture. Taking the parcellation of the human paracentral lobule by diffusion tensor imaging as an example, the paracentral lobule was first extracted from the DK atlas. The connectional architecture was then mapped with probabilistic tractography using diffusion MRI, after which, by calculating the similarity/dissimilarity between the connectivity architecture, the paracentral lobule was divided into subregions with distinguishing anatomical connectivity patterns. The stability across the population and the interhemispheric anatomic homology were evaluated to determine the final cluster number. (C) Subregional anatomical and functional connections and functional behavioral decoding. Diffusion MRI combined with tractography was used to reconstruct the major fiber bundles, while functional connectivity analysis of resting-state functional MRI was used to provide the in vivo large-scale connectivity in the human brain. We also mapped the functions to each paracentral lobule subregion via the behavioral domain and paradigm analysis using the BrainMap Database.
