- Home >> Latest News
Structural brain architectures match intrinsic functional networks and vary across domains: A study from 15000+ individuals
Na Luo1,2, Jing Sui1,2,3*, Anees Abrol4, Jiayu Chen4, Jessica A. Turner5, Eswar Damaraju4, Zening Fu4, Lingzhong Fan1,2, Dongdong Lin4, Chuanjun Zhuo6, Yong Xu7, David C. Glahn8, Amanda L. Rodrigue8, Marie T. Banich9,10, Godfrey D. Pearlson11,12,13, Vince D. Calhoun4, 14, 15 *
1Brainnetome Center and National Laboratory of Pattern Recognition, Institute of Automation, Chinese Academy of Sciences, Beijing 100190, China
2University of Chinese Academy of Sciences, Beijing 100049, China
3CAS Center for Excellence in Brain Science and Intelligence Technology, Institute of Automation, Chinese Academy of Sciences, Beijing 100190, China
4Tri-Institutional Center for Translational Research in Neuroimaging and Data Science (TReNDS): Georgia State University, Georgia Institute of Technology, and Emory University, Atlanta, GA 30303
5Department of Psychology, Neuroscience Institute, Georgia State University, Atlanta, GA, United States
6Department of Psychiatric-Neuroimaging-Genetics and Morbidity Laboratory (PNGC-Lab), Nankai University Affiliated Anding Hospital, Tianjin Mental Health Center, Tianjin, 300222, China
7Department of Psychiatry, First Clinical Medical College/ First Hospital of Shanxi Medical University, Taiyuan 030000, China
8Department of Psychiatry, Boston Children’s Hospital and Harvard Medical School, Boston, MA 02115, United States
9Department of Psychology and Neuroscience, University of Colorado Boulder, 345 UCB, Boulder, CO 80309, United States
10Institute of Cognitive Science, University of Colorado Boulder, 344 UCB, Boulder, CO 80309, United States
11Olin Neuropsychiatry Research Center, Institute of Living, Hartford, CT, United States
12Department of Psychiatry, Yale University, New Haven, CT, United States
13Department of Neuroscience, Yale University, New Haven, CT, United States
14Departments of Psychology, Computer Science, Neuroscience Institute, and Physics, Georgia State University, Atlanta, GA 30302, United States
15Department of Electrical and Computer Engineering, Georgia Institute of Technology, Atlanta, GA 30332, United States
Abstract
Brain structural networks have been shown to consistently organize in functionally meaningful architectures covering the entire brain. However, to what extent brain structural architectures match the intrinsic functional networks in different functional domains remains under explored. In this study, based on independent component analysis, we revealed 45 pairs of structural-functional (S-F) component maps, distributing across 9 functional domains, in both a discovery cohort (n=6005) and a replication cohort (UK Biobank, n=9214), providing a well-match multimodal spatial map template for public use. Further network module analysis suggested that unimodal cortical areas (e.g. somatomotor and visual networks) indicate higher S-F coherence, while heteromodal association cortices, especially the frontoparietal network (FPN), exhibit more S-F divergence. Collectively, these results suggest that the expanding and maturing brain association cortex demonstrates a higher degree of changes compared to unimodal cortex, which may lead to higher inter-individual variability and lower S-F coherence.
Keywords: intrinsic brain networks, independent component analysis (ICA), structure-function coherence, unimodal cortex, heteromodal association cortex
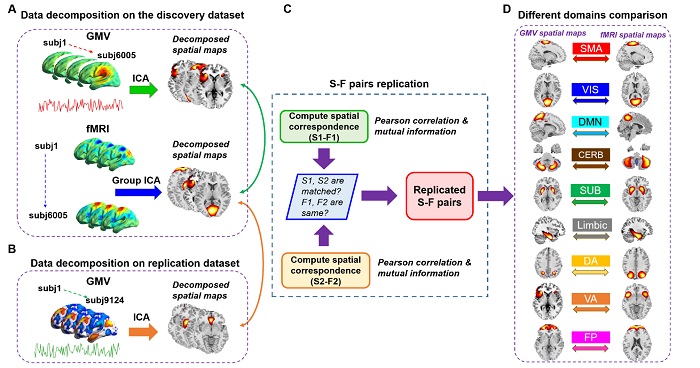 | Figure. Schematic of preprocessing and analyses pipelines. (A) Structural data and functional data were preprocessed through an automated pipeline and then decomposed using spatial ICA. We then compared the correspondence between structural and functional components using Pearson correlation and mutual information. (B) A replication dataset from UK Biobank, consisting of 9214 subjects, was preprocessed to validate the identified S-F pairs. We again applied ICA to decompose the structural replication data and measured the spatial correspondence between components in the discovery dataset and components in the replication dataset. (C) If one matched S-F pair presented a high correspondence with the same functional component, then the S-F pair in the discovery dataset was regarded as replicated. (D) We further sorted the matched pairs into 9 networks and compared S-F coherence across networks. |
